- Introduced CHASM Analyses for 10 disease cohorts:
- Machine learning method to distinguish between driver and passenger somatic missense mutations
- Driver mutations are curated from the COSMIC database
- Passenger mutations are based on background base substitution frequencies observed for the specific tumor type
- Introduced GISTIC2 analyses for each of the 13 disease studies containing Low-Pass Copy Number data (cna__illuminahiseq_dnaseqc__hms_harvard_edu)
- Mutation Assessor:
- Primary script updated such that runtime is vastly decreased for large MAF files
- New report added summarizing the functional impact of missense mutations at the gene level
- GISTIC2 updated to v2.0.19 to fix minor bugs
- Limit absurdly high CN values to +/- 1e6
- Fix line numbers reported when segment shortened to 0 markers
- Allow single- and zero-marker arms for broad analysis
- Analysis Reports:
- Increased in number to 886
- Enhanced download section of every report to note that firehose_get can be used, or Broad or TCGA sites (including URLs to each)
IMPORTANT: issues with gene annotation in GAF 3.0 have impacted mutation analyses in this run. The GAF 3.0 annotation issue has been discussed in the TCGA sequencing working group and corrective action is underway, but until then we strongly advise caution in the interpretation of mutation analyses based upon GAF 3.0. The following mutation analyses are affected in this run
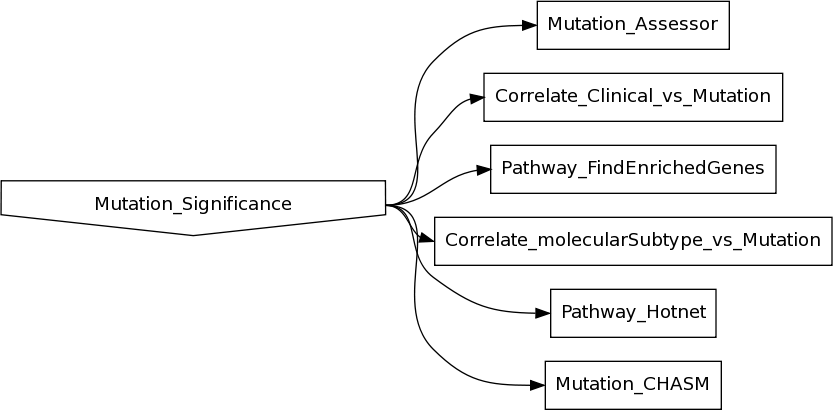
for the following 6 disease studies (which presently have MAFs with GAF 3.0 annotations deposited at the TCGA DCC): KICH, KIRP, LGG, PAAD, SKCM, THCA Rather than completely remove the affected analyses, we felt it would be valuable diagnostically to have these mutation results available for inspection. Towards that end, here is a comparison of the top 10 significant genes found by MutSig for disease studies that used a GAF 2.1 MAF in the last analyses run (2013_05_23), versus a GAF 3.0 MAF in this run: 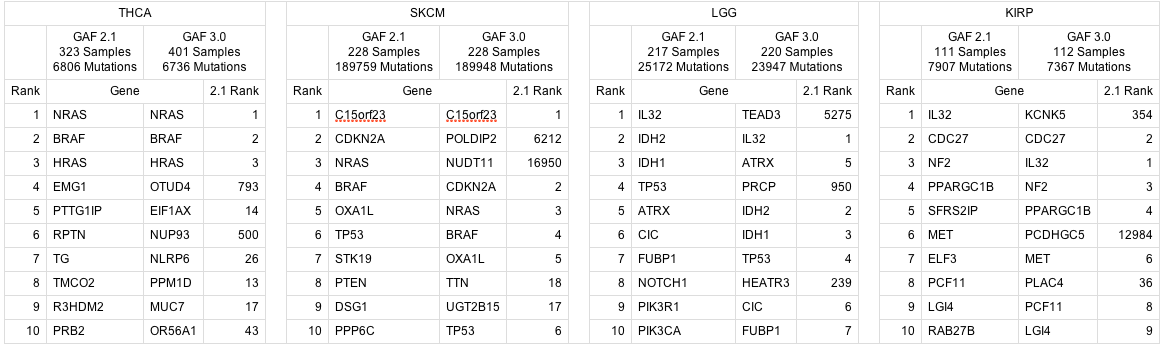
| 