Summary of sample changes (see the comprehensive samples report for more details): BCR | +15 | (11367 total) |
|---|
Clinical | +106 | (11164 total) |
|---|
CN | -2 | (10985 total) |
|---|
LowP | +120 | (1211 total) |
|---|
MAF | +48 | (6786 total) |
|---|
Methylation | +16 | (10971 total) |
|---|
miRSeq | -4 | (10154 total) |
|---|
mRNASeq | +8 | (10103 total) |
|---|
RPPA | +650 | (6802 total) |
|---|
Extensive improvements to clinical data, in that Clinical_Pick_Tier1 archives now bundle 2 forms of values: Entire set of TCGA CDEs, verbatim (in new All_CDES.txt file): adding over 700 additional clinical parameters In addition to the CDE subset normalized by Firehose for downstream analyses (in <cohort>.clin.merged.picked.txt file) - For example, to date the ACC picked file has contained less than 20 CDEs while
All_CDEs.txt now contains more than 100. - Followup values are merged, when available, to yield the most up-to-date values per CDE
- Corrected problem wherein some True/False values for
regimen_indication CDE were erroneously swapped - Created an interactive table CDEs, which on a single page shows exactly what CDEs are selected for analyses in Firehose for all disease cohorts. Updated the FireBrowse clinical samples API to reference this new CDE table
- Enhanced
Merge_Clinical pipeline to leverage auxiliary CDEs when available (COAD, READ, ESCA): for all primary CDEs that also have a value in the aux CDE file (e.g. MSI), we now replace the primary value if it is NA and the aux value is not NA
- Extensive improvements to TCGA mutation data:
- New MAF for Diffuse Large B-cell Lymphoma (DLBC, 48 mutation samples)
- Oncotator now included in Firehose mutation pipelines, to standardize TCGA MAFs to a common format:
- This substantially improves the consistency and utility of TCGA MAFs
- hg18 MAFs lifted over to hg19
- All MAFs re-annotated against Gencode v19
- Oft-requested custom columns, such as amino acid change, now present in all MAFs
- Oncotated MAFs are available in 2 pipeline output archives
- Mutation_Packager_Oncotated_Calls
- Mutation_Packager_Oncotated_Raw_Calls
reflecting the separation of MAFs into two sets (raw/automated and curated, per the Spring 2015 analysis run)
- Extensive improvements to RPPA data, fostering more robust automated processing and downstream analysis:
- Merge_protein_exp__mda_rppa_core__mdanderson_org__Level_3__protein_normalization__data
- Validation and animal source suffixes now stripped off of antibody name to account for several new batches that no longer include them
- Now returns a union of antibody names from all samples, rather than failing when all samples don't have the exact same antibodies
- This does not fix the below RPPA issues when different names for the same antibodies are used (e.g. Acetyl-a-Tubulin-Lys40/Acetyl-a-Tubulin(Lys40), ARHI/DIRAS3) - in these cases each antibody name will appear on separate lines of the merged files until a fix is made at MDACC.
- This enables RPPA analysis for aggregate cohorts such as STES, KIPAN, and GBMLGG
- RPPA_AnnotateWithGene: normalizes the antibody reference files provided by MDACC into a two-column tsv with standardized header, gene name, and antibody name (stripped of suffixes). This file is now provided in the archive.
- FireBrowse v1.1.17 beta:
- Ingest this August 2015 data run:
API through which Firehose-picked and normalized clinical data are accessible has been renamed to Samples/Clinical_FH Verbatim TCGA clinical data may be accessed through the new Samples/Clincal API A CDE will be reflected in either API only when it has a value other than NA for at least 1 patient case in any disease cohort. - For backward compatibility, the Samples/ClinicalTier1 remains available (as a synonym for Samples/Clinical_FH)
- fbget Python and UNIX CLI bindings have been suitably updated
- Which makes it extremely easy to determine for what patients and cohorts any given CDE is defined: e.g.
fbget clinical cde=gleason_score
will show each patient case with a valid gleason_score, across all cohorts
- viewGene: now enforces rendering of at most 1 gene per Submit; if multiple genes are given the first is selected
- iCoMut:
- Popup tooltips for mutation panels now show total # of mutations AND fractional % of a given type (e.g. missense)
- New search feature, enabling one to see into which clusters/panels/etc a given patient (or set of patients) falls
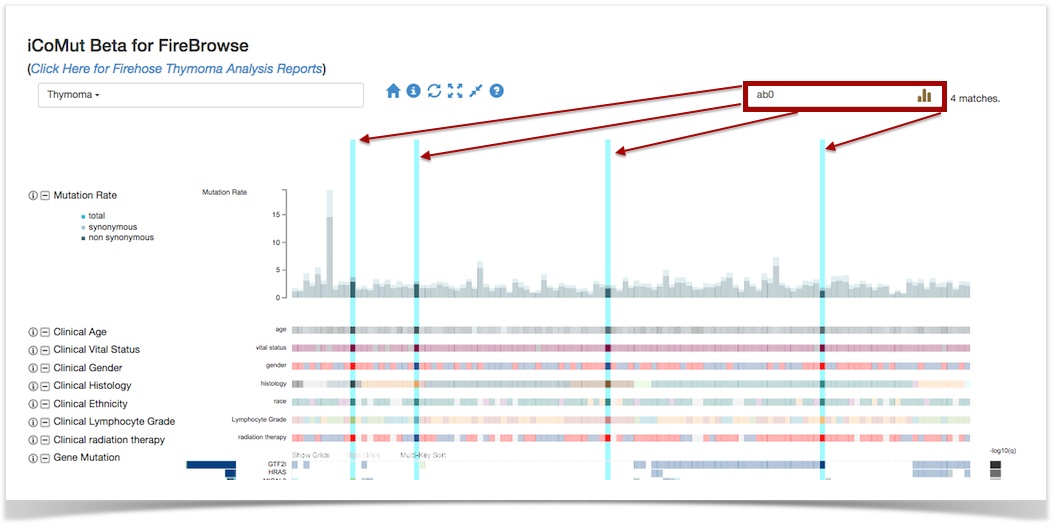
- After extensive testing, upgraded backend database from Mongo2.x to Mongo3.x: v3 dramatically reduces the memory footprint and data storage sizes, which yields greater performance and also clears considerable breathing room to add more data APIs (e.g. for methylation, RPPA, etc) as well as AWG-specific databases
- Corrected item # 9 from Spring 2015 data run: missing RPKM aliqouts in COAD
- Issues:
- RPPA issues due to changes in new data file row names: these have been previously reported to MDACC.
- KIRP: ARHI-M-E -> DIRAS3-M-E (batch 2.0.0)
- LGG: Acetyl-a-Tubulin-Lys40-R-C -> Acetyl-a-Tubulin(Lys40)-R-C (batch 2.0.0)
- PAAD: DIRAS3-M-E -> ARHI-M-E (batch 1.2.0)
- STES: ARHI-M-E -> DIRAS3-M-E (batch 2.0.0)
- RPPA issue with KIRC antibody file:
Gene names were missing for 15 antibodies. The gene names were located in an online supplement file, and manually added: CA9 | CA9 | SDHB | Complex-II_subunit30 | GYG1 | GYG-Glycogenin1 | GYS1 | GYS | GYS1 | GYS_pS641 | HIF1A | HIF-1_alpha | LDHA | LDHA | LDHB | LDHB | MTCO2 | Mitochondria | ATP5A1 | Oxphos-complex-V_subunitb | PKM2 | PKM2 | PYGB | PYGB | PYGB | PYGB-AB2 | PYGL | PYGL | PYGM | PYGM |
|